DAPRed - Autophagy Detection

Autophagy (Autophagosome) Detection
-
Product codeD677 DAPRed - Autophagy Detection
| Unit size | Price | Item Code |
|---|---|---|
| 5 nmol | D677-10 |
Detection Principle
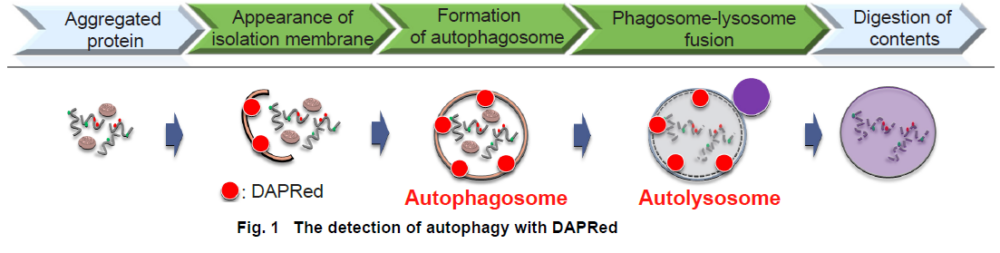
Autophagy is an intracellular degradation system, where dysfunctional proteins and organelles are degraded. In this process, aggregated dysfunctional proteins are surrounded by the double membrane to form an autophagosome.
The small fluorescent molecule DAPRed is used to be detect autophagosomes and autolysosomes. The mechanism has been suggested to be that the dye is incorporated into the autophagosome during double-membrane formation via structural features, and then emits fluorescence under hydrophobic conditions. The utility of DAPRed is conferred by its molecular properties: it is permeable to cells, has no requirement for transfection, and enables live cell imaging with fluorescence microscopy. For monitoring autolysosomes, DALGreen (D675) is recommend because it enables the detection of phagosome-lysosome fusion.
Manual
Technical info
Gene transfection is not necessary. You only need to add the reagent to your cell sample, and you can get a fluorescent image.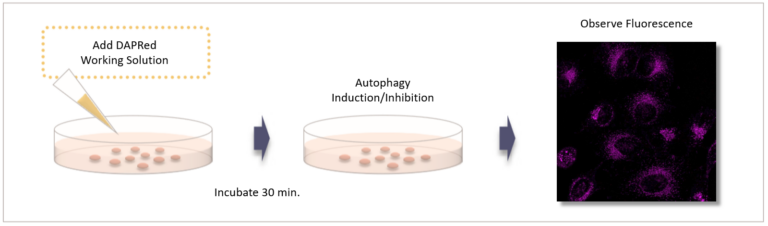
Co-staining of DAPRed and DALGreen
HeLa cells were stained with DAPRed, which is a dye-stained autophagosome, and DALGreen, which is a dye-stained autolysosome. Autophagy was then induced by starvation.
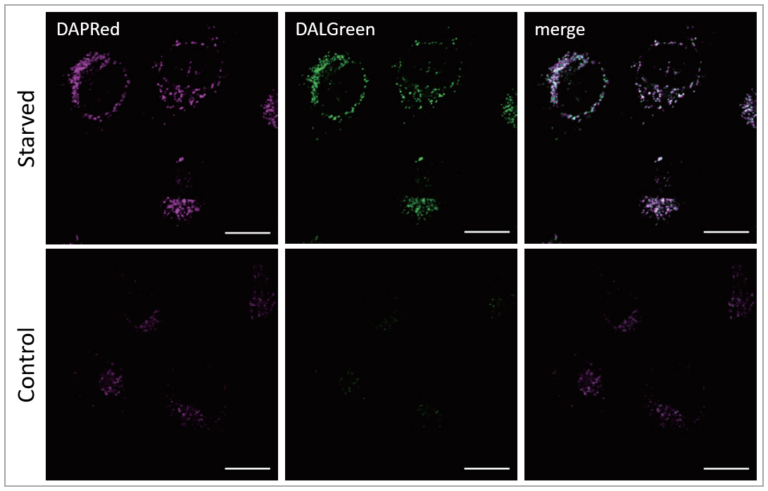
Result:
The fluorescence of DAPRed and DALGreen was stronger in starved HeLa cell culture.
Detection condition:
DAPRed: Ex. 561 nm/Em. 600-700 nm
DALGreen: Ex. 488 nm/Em. 500-563 nm
Scale bar: 20 μm
Autophagy Induction:
After staining with DAPRed and DALGreen, HeLa cell was incubated with culture medium or amino acid-free medium for 5 hours.
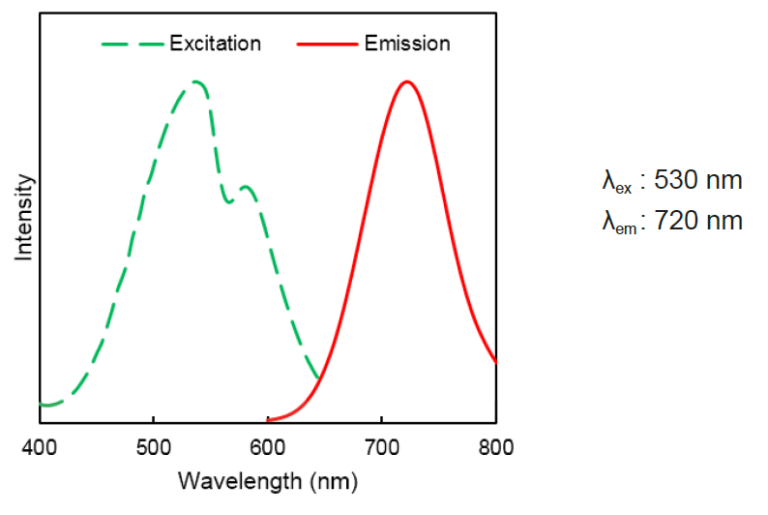
DAPRed Excitation/Emission
Related Product Information
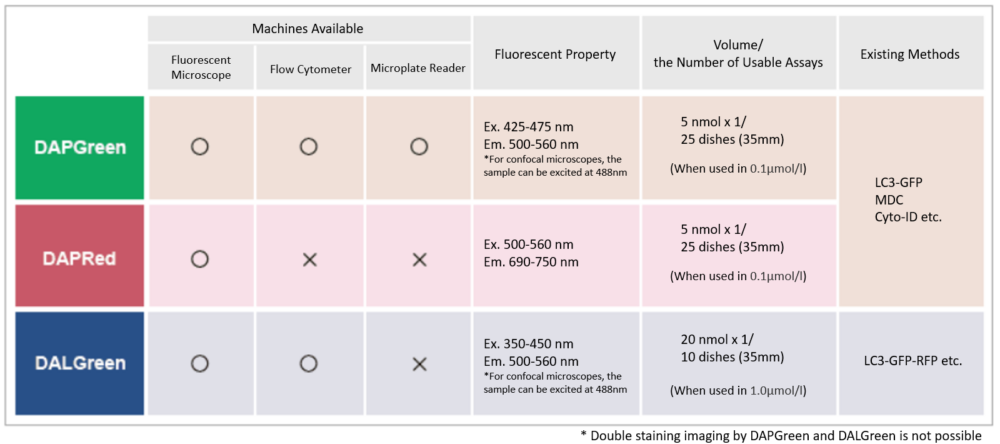
Detection is possible with a fluorescent microscope, a flow cytometer, and a microplate reader using DAPGreen [#D676] and DAPRed. DALGreen [#D675] detects autolysosome. After a lysosome fuses with the autophagosome, the environment in the autolysosome becomes acidic. DALGreen fluoresces stronger as acidity increases. DALGreen can be applied in two methods (a fluorescent microscope and a flow cytometer). Please select the most suitable method depending on your equipment.
References
| No. | Sample | Instrument | Reference (Link) |
|---|---|---|---|
| 1) | Cell (HUVEC) |
Fluorescence microscope | X. Chen, X. Yan, J. Liu and L. Zhang, " Chaiqi decoction ameliorates vascular endothelial injury in metabolic syndrome by upregulating autophagy.", Am. J. Transl. Res., 2020,12(9), 4902. |
| 2) | Cell (HeLa) |
Fluorescence microscope | H. Fang , S. Geng, M. Hao, Q. Chen, M. Liu, C. Liu, Z. Tian, C. Wang, T. Takebe, J-L Guan, Y. Chen, Z. Guo, W. He and J. Diao, "Simultaneous Zn2+ tracking in multiple organelles using super-resolution morphology-correlated organelle identification in living cells ", Nat Commun, 2021, 12(1), 109. 10.1016/j.envpol.2019.07.105. |
| 3) | Cell (HCT116; HCT8) |
Fluorescence microscope | H. Sun, R. Wang, Y. Liu, H. Mei and X. Liu , "USP11 induce resistance to 5-Fluorouracil in Colorectal Cancer through activating autophagy by stabilizing VCP", J Cancer , 2021, 12(8), 2317. |
| 4) | Cell (MEF) |
Fluorescence microscope | M. Yagi, T. Toshima, R. Amamoto, Y. Do, H. Hirai, D. Setoyama, D. Kang and T. Uchiumi, "Mitochondrial translation deficiency impairs NAD+-mediated lysosomal acidification", EMBO J, 2021, doi:10.15252/embj.2020105268. |
| 5) | Cell (SH-SY5Y) |
Fluorescence microscope | Chang-ki Oh, Nima Dolatabadi, Piotr Cieplak, Maria T. Diaz-Meco, Jorge Moscat, John P. Nolan, Tomohiro Nakamura and Stuart A. Lipton, "S-Nitrosylation of p62 Inhibits Autophagic Flux to Promote α-Synuclein Secretion and Spread in Parkinson’s Disease and Lewy Body Dementia", J. Neurosci., 2022, doi:10.1523/JNEUROSCI.1508-21.2022. |
Q & A
-
Q
How stable is DAPRed working solution?
-
A
DAPRed working solution cannot be stored and should be used up on the day it is prepared.
-
Q
How stable is DAPRed DMSO stock solution?
-
A
The stock solution should be stored at -20°C in a light-shielded condition and is stable for 1 month after preparation.
For long storage, it is recommended to divide the solution into smaller portions.
-
Q
What excitation and emission conditions do you recommend?
-
A
Excitation: 500 - 560 nm
Emission: 690 - 750 nm
-
Q
How do I decide the optimal concentration of DAPRed?
-
A
If the reagent concentration is too high or too low, it may be difficult to tell the difference between autophagy induction and uninduced control.
It is recommended to consider the reagent concentration concerning the following information.The optimal concentration of DAPRed depends on the cell type.
Please consider increasing the concentration of DAPRed in steps of several points starting from a low concentration (0.05 - 0.4 μmol/l is the approximate range).(Reference)
We have studied the optimal concentration of DAPRed for each cell type in HeLa, HepG2, and CHO cells.
DAPRed was stained at the following concentrations and cultured in an amino acid-free medium to induce autophagy.As a result, differences from the control were observed under the following concentration conditions highlighted in red.
Cell types Tested concentration HeLa 0.4 µmol/l, 0.2 µmol/l, 0.1 µmol/l, 0.05 µmol/l HepG2 0.4 µmol/l, 0.2 µmol/l, 0.1 µmol/l, 0.05 µmol/l CHO 0.4 µmol/l, 0.2 µmol/l, 0.1 µmol/l, 0.05 µmol/l HeLa
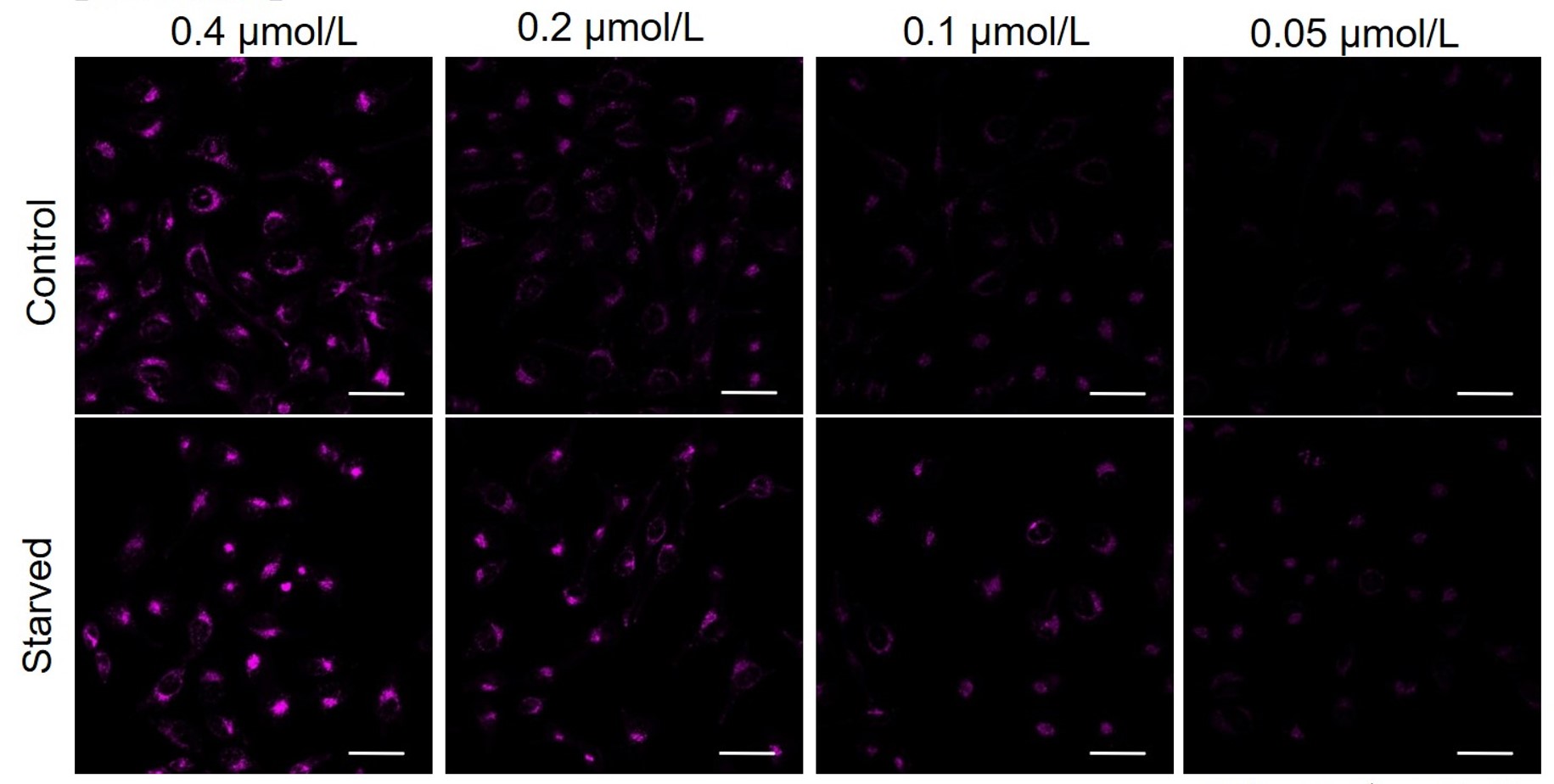
HepG2
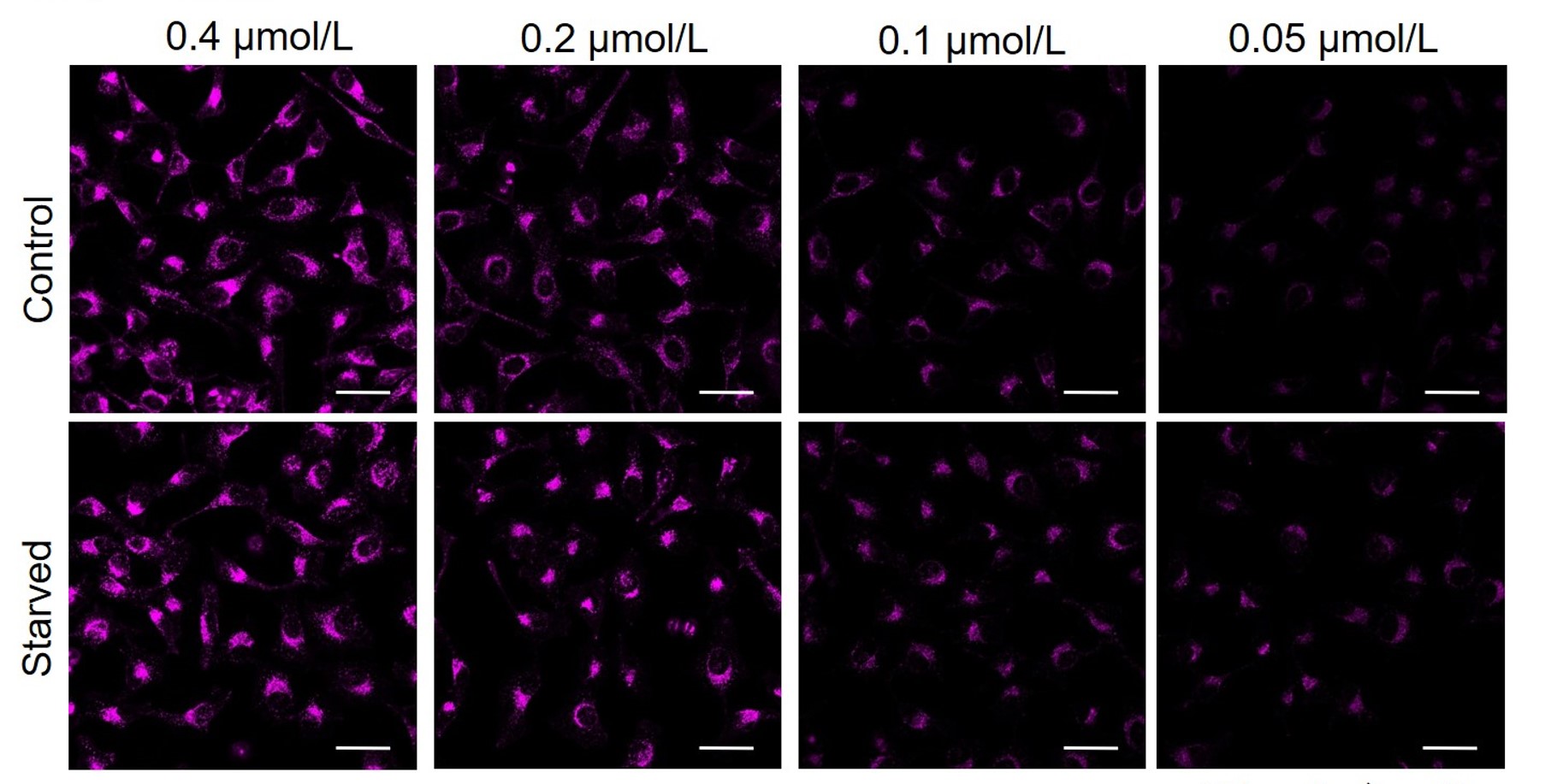
CHO
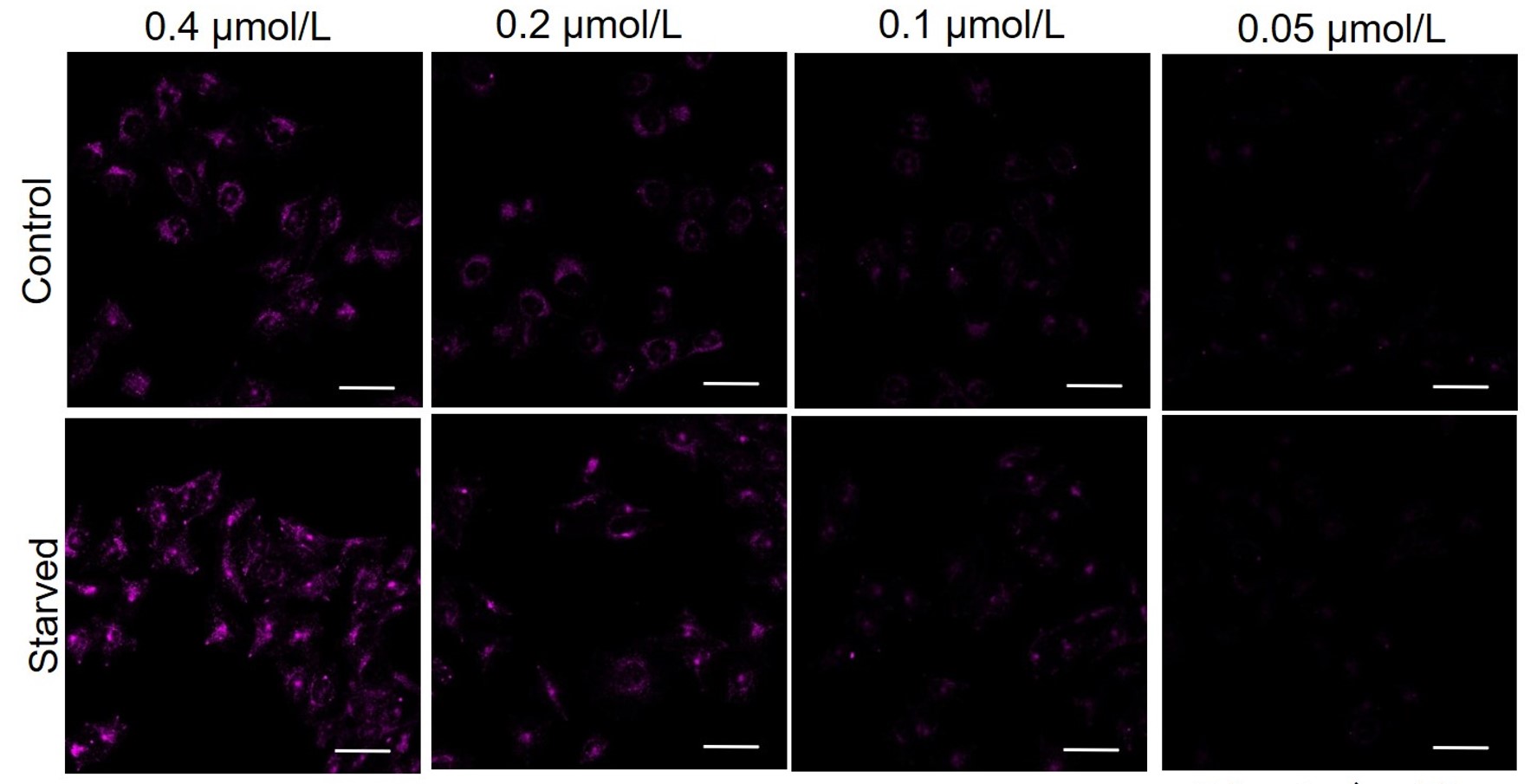
All Imaging Conditions:
DAPRed: Ex. 561 nm/Em. 600-700 nm
Scale bar: 50 μm
Handling and storage condition
| Appearance: | Reddish brown solid |
|---|---|
| Purity (HPLC): | ≧ 93.0 % |
| 0-5°C, Protect from light |


















