JC-1 MitoMP Detection Kit

Mitochondrial Membrane Potential Detection
-
Product codeMT09 JC-1 MitoMP Detection Kit
| Unit size | Price | Item Code |
|---|---|---|
| 1 set | MT09-10 |
| 1 set | JC-1 Dye Imaging Buffer (10x) |
100 nmol x1 11 ml x1 |
|---|
Description
Mitochondria synthesize ATP using oxygen to produce necessary energy for living cells. Lowering of mitochondrial activity and dysfunction are known to be closely related to cancer, aging, and neurodegenerative diseases such as Alzheimer’s and Parkinson’s diseases. Mitochondrial membrane potential is a parameter used to measure with mitochondrial condition.
Manual
Technical info
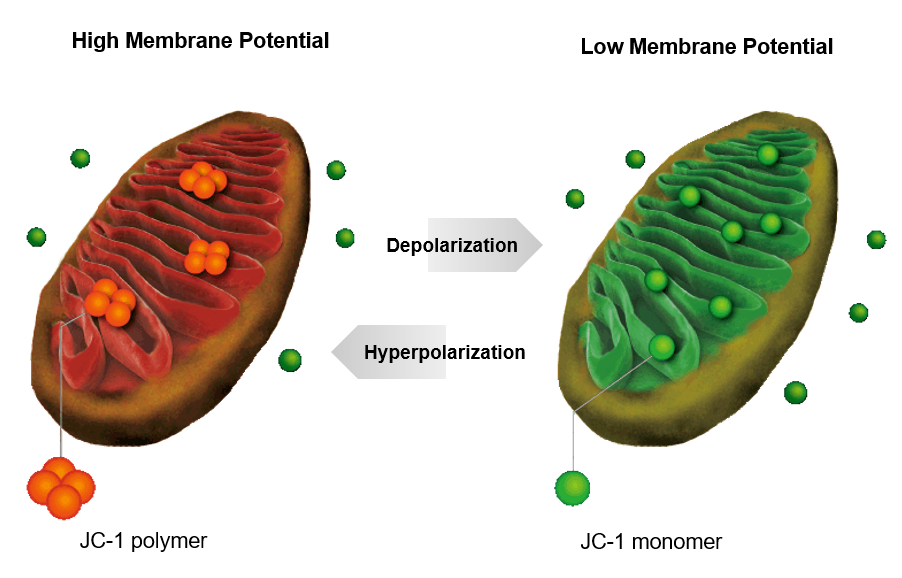
JC-1 forms aggregate (in healthy mitochondria) with red fluorescence. As membrane potential decreases, JC-1 becomes monomers, which shows in green fluorescence. The change in ratio of red to green fluorescence is used as a indicator of mitochondrial condition.
Easy to Use
|
Easy to dissolve JC-1 has been difficult to dissolve, but this kit solves the problem!
|
Detect by Several Equipments Please refer to Data: Induced Apoptosis for experimental examples
|
Imaging Buffer Included |
Procedure
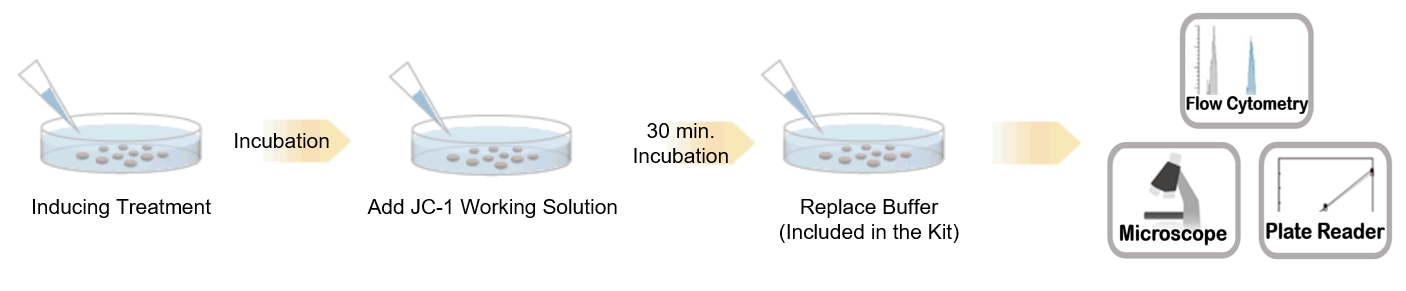
Comparison of reagents for mitochondrial membrane potential detection
| Features | Sensitivity | Fixation | Monitoring | Fluorescence change (upon loss of mitochondrial membrane potential) |
Detection |
|
|---|---|---|---|---|---|---|
|
JC-1 |
Recomended for starting-up |
✓ |
|
|
Color change from red to green |
|
|
Recommended for more detailed analysis |
✓ (High) |
✓ |
✓ |
Decrease in fluorescence intensity |
530-560 nm / 570-640 nm |
|
|
TMRE |
Widely used |
✓ (High) |
|
|
Decrease in fluorescence intensity |
530-560 nm / 570-640 nm |
Data: Depolarization
HeLa cells treated with depolarizing reagent, carbonylcyanide-p-trifluoromethoxyphenylhydrazone (FCCP) were stained with JC-1 MitoMP Detection Kit. Red fluorescence indicates normal membrane potential or health mitochondria. Untreated cells showed red fluorescence, while FCCP treated cells showed little red fluorescence.
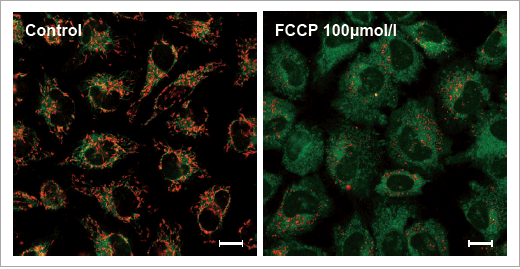
|
Data: Induced Apoptosis
Jurkat cells treated by apoptosis inducing reagent, Staurosporine, were stained with JC-1 MitoMP Detection Kit. Procedures for these data can be found in the Technical Manual.
[Fluorescence Microscope]
Fluorescence imaging of mitochondrial membrane potential in Jurkat cells
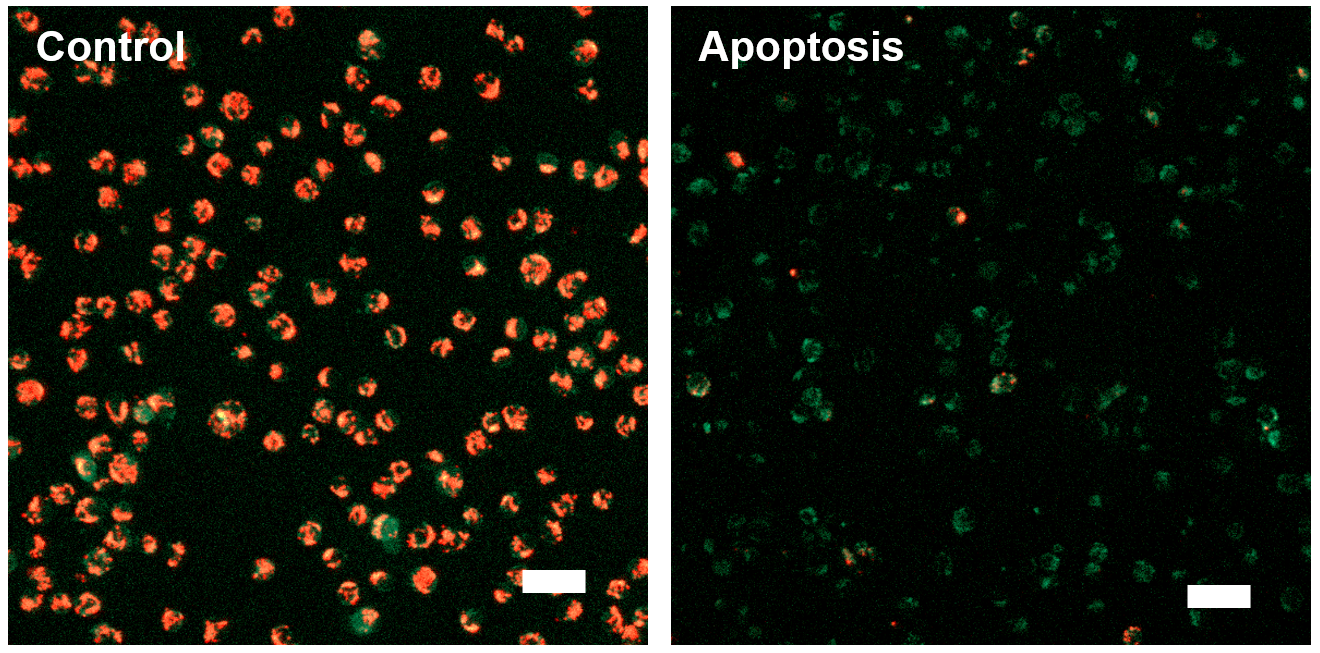
|
[Flow Cytometry]
Flow cytometric analysis of mitochondrial membrane potential in Jurkat cells
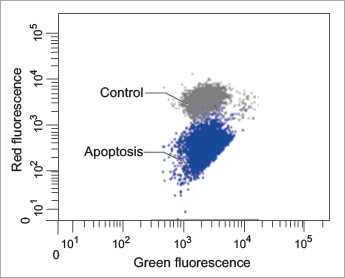
|
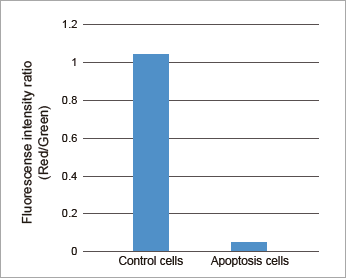
Plate Reader
Fluorescence intensity ratio of mitochondrial membrane potential in Jurkat cells
|
Required amount of Imaging Buffer solution by vessel type

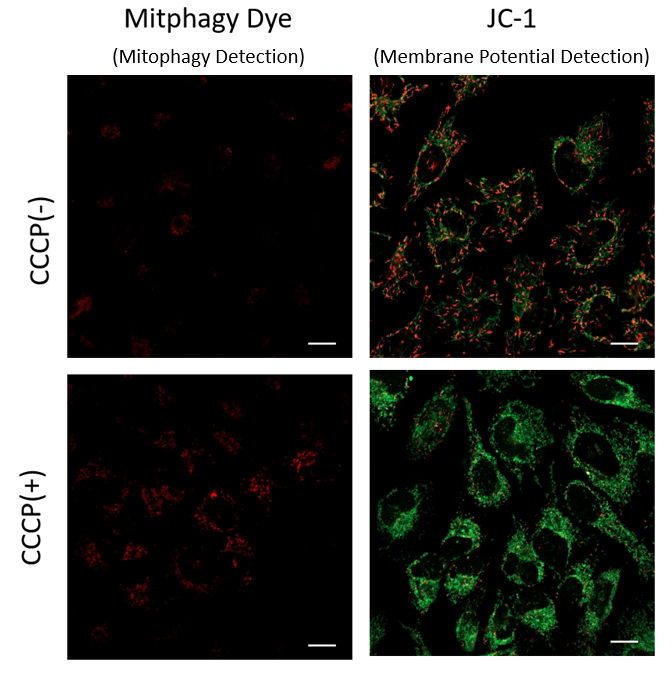
Mitophagy Induction and Detection of Mitochondrial Membrane Potential Changes
Mitochondrial condition in the carbonyl cyanide m-chlorophenyl hydrazine (CCCP) treated Parkin-expressing HeLa cells was compared with untreated cells using Mitophagy Detection Kit (MD01, MT02) and JC-1 MitoMP Detection Kit (MT09).
Result:
Mitophagy was not detected in untreated cells and the membrane potential was normal. However, reduction of membrane potential and mitophagy were observed in treated cells.
|
Detecting Condition
[Mitophagy Detection]
Ex: 561 nm, Em: 570-700 nm
[Mitochondrial Membrane Potential Detection]
Green Ex: 488 nm, Em: 500-550 nm
Red Ex: 561 nm, Em: 560-610 nm
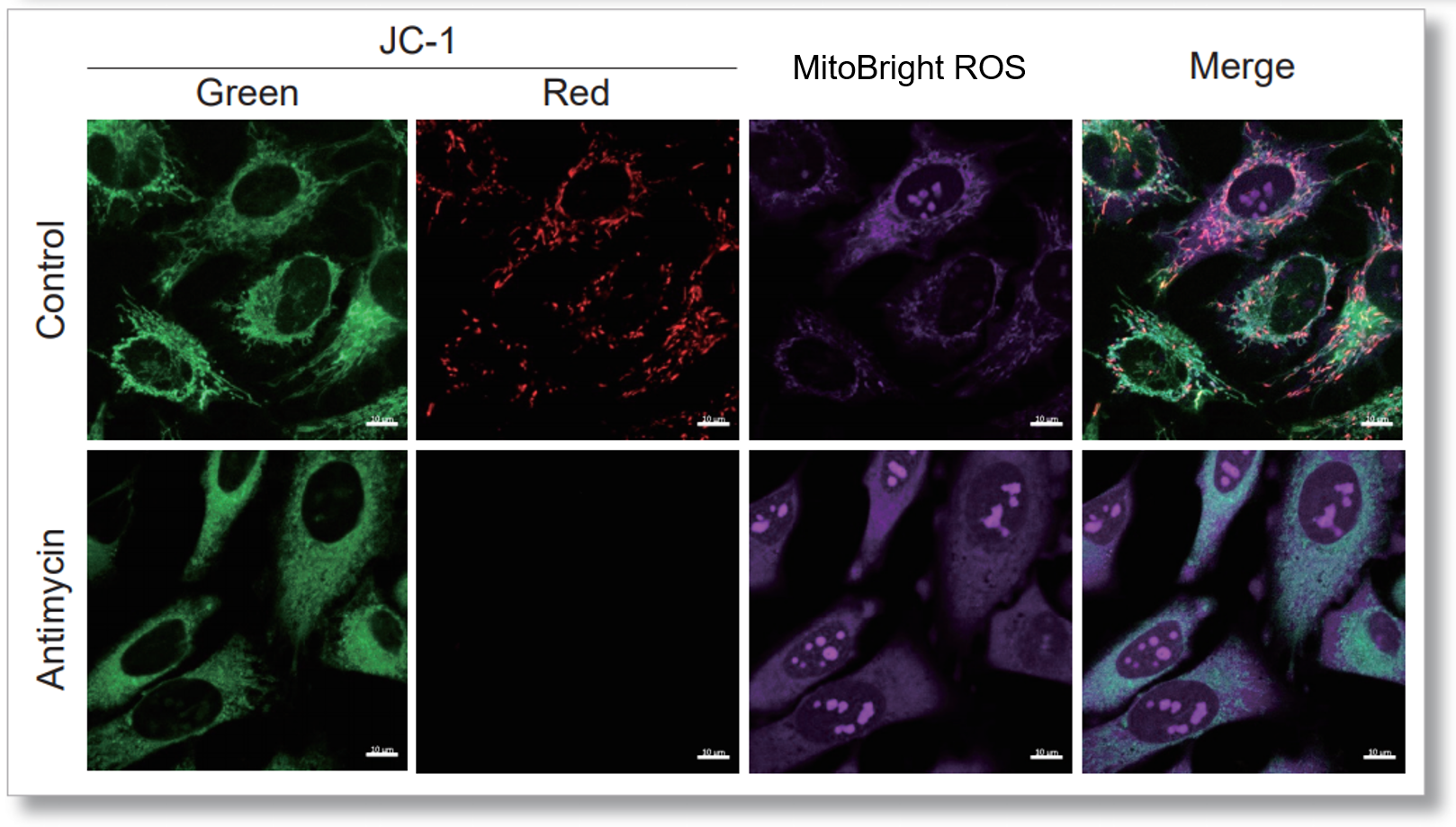
Application Data: Simultaneously evaluation of mitochondrial superoxide and membrane potential
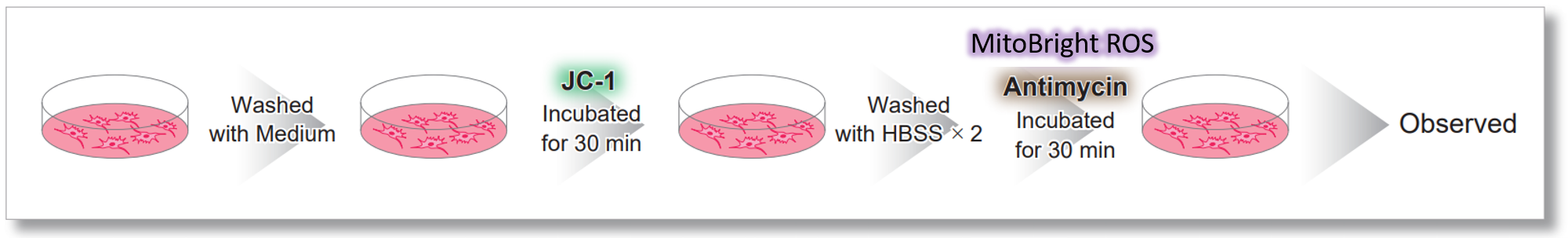
After HeLa cells were washed with HBSS, co-stained with MitoBright ROS Deep Red and mitochondrial membrane potential staining dye (JC-1: code MT09), and the generated mitochondrial ROS and membrane potential were observed simultaneously. As a result, the decrease in mitochondrial membrane potential and the generation of mitochondrial ROS are simultaneously observed.
<General protocol of JC-1>


<Imaging Conditions>(Confocal microscopy)
JC-1: Green Ex = 488, Em = 490-520 nm, Red: Ex = 561, Em = 560-600 nm
MitoBright ROS :Ex = 633 nm, Em = 640-700 nm
Scale bar: 10 μm

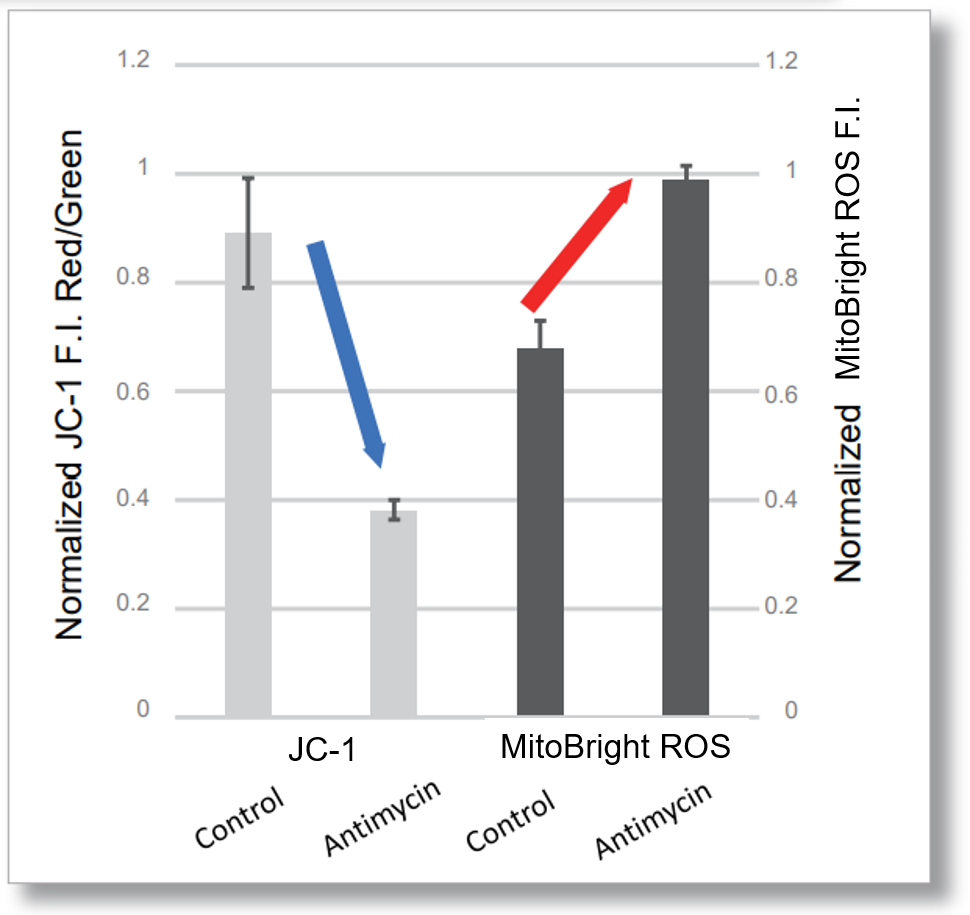
<Examination Conditions>(Plate Reader)Tecan, Infinite M200 Pro
JC-1: Green Ex=480-490 nm, Em=525-545 nm; Red: Ex= 530-540 nm, Em=585-605 nm
MitoBright ROS: Ex=545-555 nm, Em = 665-685 nm
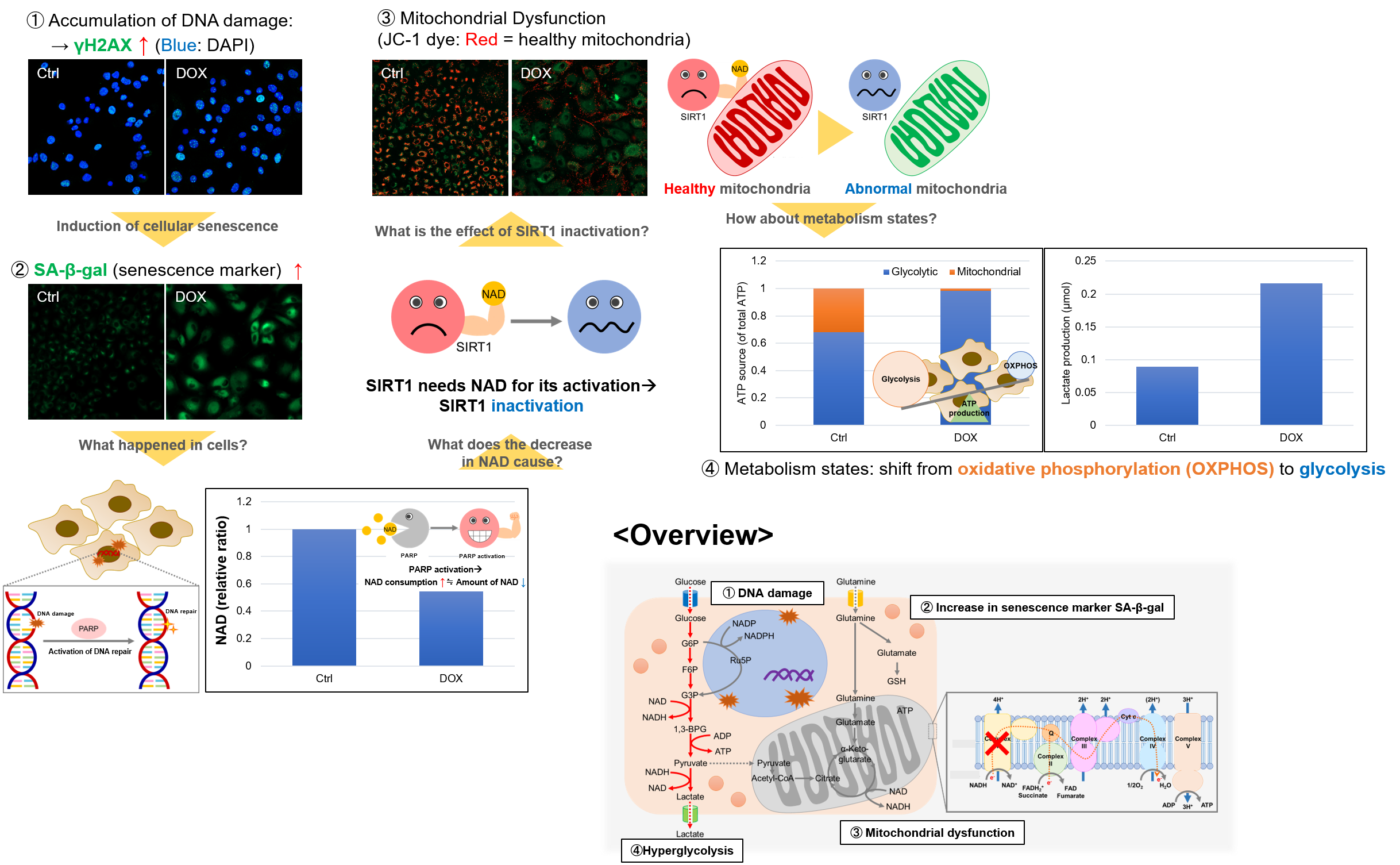
Experiment Example: Cellular Senescence with Metabolic shift
NAD(+) levels decline during the aging process, causing defects in nuclear and mitochondrial functions and resulting in many age-associated pathologies*. Here, we try to redemonstrate this phenomenon in the doxorubicin (DOX)-induced cellular senescence model with a comprehensive analysis of our products.
*S. Imai, et al., Trends Cell Biol, 2014, 24, 464-471
Related Products
| Category | Product | Unit | SKU |
|---|---|---|---|
|
Cellular Metabolism |
100 tests |
N509-10 |
|
| Lactate Assay Kit-WST | 50 / 200 tests | L256-10 / L256-20 | |
| Glycolysis/OXPHOS Assay Kit NEW | 50 tests | G270-10 | |
|
Cellular Senescence Detection |
1 / 3 / 10 plates |
SG04-01 / SG04-03 / SG04-10 |
|
|
Mitochondrial Membrane Potential |
10 set |
MT09-10 |
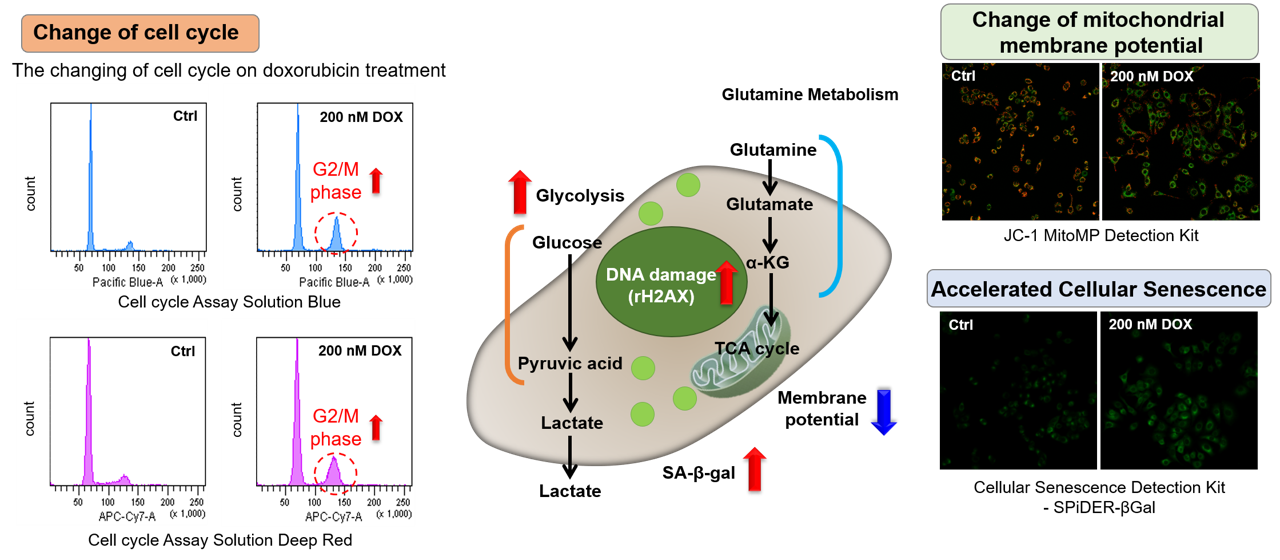
Association between cellular senescence and cell cycle
Doxorubicin (DOX) acts to inhibit cell proliferation during G2/M phases of the cell cycle and induces cellular senescence. After adding DOX to A549 cells, higher histogram peaks for the G2/M phase (Cell Cycle Assay Solution Blue and Deep Red), induces cellular senescence (Cellular Senescence Detection Kit - SPiDER-βGal), and the differences in mitochondrial membrane potential (JC-1 MitoMP Detection Kit) were observed.
References
| No. | Sample Type | Instrument | Reference |
|---|---|---|---|
| 1 | Cell: A549 |
Microscope | K. Li, S. Sun, L. Xiao and Z. Zhang, "Bioactivity-guided fractionation of Helicteres angustifolia L. extract and its molecular evidence for tumor suppression", Front Cell Dev Biol.,2023, doi: 10.3389/fcell.2023.1157172. |
| 2 | Cell: A549 |
Flow Cytometer | C. N. D’Alessandro-Gabazza, T. Yasuma, T. Kobayashi, M. Toda1, A. M. Abdel-Hamid, H. Fujimoto, O. Hataji, H. Nakahara, A. Takeshita, K. Nishihama, T. Okano, H. Saiki, Y. Okano, A. Tomaru, V. F. D’Alessandro, M. Shiraishi, A. Mizoguchi, R. Ono, J. Ohtsuka, M. Fukumura, T. Nosaka, X. Mi, D. Shukla, K. Kataoka, Y. Kondoh, M. Hirose, T. Arai, Y. Inoue, Y. Yano, R. I. Mackie, I. Cann and E. C. Gabazza, "Inhibition of lung microbiota-derived proapoptotic peptides ameliorates acute exacerbation of pulmonary fibrosis", Nat. Comm., 2022, doi:10.1038/s41467-022-29064-3. |
| 3 | Cell: A549, HeLa |
Plate reader | J. Yang, L. Liu, Y. Oda, K. Wada, M. Ago, S. Matsuda, M. Hattori, T. Goto, Y. Kawashima, Y. Matsuzaki and T. Taketani,"Highly-purified rapidly expanding clones, RECs, are superior for functional-mitochondrial transfer", Stem Cell Res Ther., 2023, doi: 10.1186/s13287-023-03274-y. |
| 4 | Cell: ALM |
Plate reader | T. Nechiporuk, S.E. Kurtz, O. Nikolova, T. Liu, C.L. Jones, A. D. Alessandro, R. C. Hill, A. Almeida, S. K. Joshi, M. Rosenberg, C. E. Tognon, A. V. Danilov, B. J. Druker, B. H. Chang, S. K McWeeney and J. W. Tyner , "The TP53 Apoptotic Network Is a Primary Mediator of Resistance to BCL2 Inhibition in AML Cells.", Cancer Discov, 2019, 9, |
| 5 | Cell: ARPE-19 |
Flow Cytometer/ Microscope |
J. Hamuro, T. Yamashita, Y. Otsuki, N. Hiramoto, M. Adachi, T. Miyatani, H. Tanaka, M. Ueno, S. Kinoshita and C. Sotozono,"Spatiotemporal Coordination of RPE Cell Quality by Extracellular Vesicle miR-494-3p Via Competitive Interplays With SIRT3 or PTEN", Invest Ophthalmol Vis Sci., 2023, doi: 10.1167/iovs.64.5.9. |
| 6 | Cell: ARPE-19 |
Microscope | J. H. Quan, F. F. Gao, H. A. Ismail, J. M. Yuk, G. H. Cha, J. Q. Chu and Y. H. Lee, "Silver Nanoparticle-Induced Apoptosis in ARPE-19 Cells Is Inhibited by Toxoplasma gondii Pre-Infection Through Suppression of NOX4-Dependent ROS Generation", Int J Nanomedicine., 2020, 15, 3695–3716. |
| 7 | Cell: C2C12, myocytes |
- | Z. Jing, T. Iba, H. Naito, P. Xu, J.I. Morishige, N. Nagata, H. Okubo and H.Ando ,"L-carnitine prevents lenvatinib-induced muscle toxicity without impairment of the anti-angiogenic efficacy", Front Pharmacol., 2023, doi: 10.3389/fphar.2023.1182788. |
| 8 | Cell: C2C12, 3T3L1 |
Plate reader | M. Kurano, K. Tsukamoto, T. Shimizu, H. Kassai, K. Nakao, A. Aiba, M. Hara and Yatomi , "Protection Against Insulin Resistance by Apolipoprotein M/Sphingosine 1-Phosphate ", Diabetes, 2020, DOI: 10.2337/db19-0811. |
| 9 | Cell: Colon 26 |
Microscope | B. Uranbileg, M. Kurano, K. Kano, E. Sakai, J. Arita, K. Hasegawa, T. Nishikawa, S. Ishihara, H. Yamashita, Y. Seto, H. Ikeda, J. Aoki and Y. Yatomi,"Sphingosine 1‐phosphate lyase facilitates cancer progression through converting sphingolipids to glycerophospholipids", Clin Transl Med., 2022, doi: 10.1002/ctm2.1056. |
| 10 | Tissue: Frozen heart slides |
Microscope | W. Yu, Y. Hu, Z. Liu, K. Guo, D. Ma, M. Peng, Y. Wang, J. Zhang, X. Zhang, P. Wang, J. Zhang, P. Liu and J. Lu,"Sorting nexin 3 exacerbates doxorubicin-induced cardiomyopathy via regulation of TFRC-dependent ferroptosis", Acta Pharmaceutica Sinica B., 2023, doi: https://doi.org/10.1016/j.apsb.2023.08.016. |
| 11 | Cell: HCE |
Microscope | T. Yamashita, K. Asada, M. Ueno, N. Hiramoto, T. Fujita, M. Toda, C. Sotozono, S. Kinoshita and J. Hamuro,"Cellular interplay through extracellular vesicle miR-184 alleviates corneal endothelium degeneration", Ophthalmol Sci., 2022, doi: 10.1016/j.xops.2022.100212. |
| 12 | Cell: HCE |
Microscope | M. Ueno, K Yoshii, T. Yamashita, K. Sonomura, K. Asada, E. Ito, T. Fujita, C. Sotozono, S. Kinoshita and J. Hamuro,"The Interplay Between Metabolites and MicroRNAs in Aqueous Humor to Coordinate Corneal Endothelium Integrity", Ophthalmol Sci., 2023, doi: 10.1016/j.xops.2023.100299. |
| 13 | Cell: HCE-T |
- | W. Otsu, T. Yako, E. Sugisawa, S. Nakamura, H. Tsusaki, N. Umigai, M. Shimazawa and H. Hara,"Crocetin protects against mitochondrial damage induced by UV-A irradiation in corneal epithelial cell line HCE-T cells", J Pharmacol Sci., 2022, doi: 10.1016/j.jphs.2022.10.005. |
| 14 | Cell: HCE-T |
Microscope | K. Ishida, T. Yako, M. Tanaka, W. Otsu, S. Nakamura, M. Shimazawa, H. Tsusaki and H. Hara,"Free-radical scavenger NSP-116 protects the corneal epithelium against UV-A and blue led light exposure", Biol Pharm Bull., 2021, doi: 10.1248/bpb.b21-00017. |
| 15 | Cell: HepG |
Microscope/ Spectrophotometer |
M. Ikura, K. Furuya, T. Matsuda and T. Ikura,"Impact of Nuclear De Novo NAD+ Synthesis via Histone Dynamics on DNA Repair during Cellular Senescence To Prevent Tumorigenesis", Mol Cell Biol., 2022, doi: 10.1128/mcb.00379-22. |
| 16 | Cell: hiPSCs, Neurons |
Microscope | T. Hara, M. Toyoshima, Y. Hisano, S. Balan, Y. Iwayama, H. Aono,Y. Futamura, H. Osada, Y. Owada and T. Yoshikawa,"Glyoxalase I disruption and external carbonyl stress impair mitochondrial function in human induced pluripotent stem cells and derived neurons", Translational Psychiatry., 2021, doi: 10.1038/s41398-021-01392-w. |
| 17 | Cell: HSCs |
Microscope | Y. Su, S. Lu, C. Hou, K. Ren, M. Wang, X. Liu, S. Zhao and X. Liu ,"Mitigation of liver fibrosis via hepatic stellate cells mitochondrial apoptosis induced by metformin", International Immunopharmacology., 2022, doi: 10.1016/j.intimp.2022.108683. |
| 18 | Cell: HUVECs |
Microscope | D. Ueno, K. Ikeda, E. Yamazaki, A. Katayama, R. Urata and S. Matoba ,"Spermidine improves angiogenic capacity of senescent endothelial cells, and enhances ischemia-induced neovascularization in aged mice", Sci Rep., 2023, doi: 10.1038/s41598-023-35447-3. |
| 19 | Cell: KYSE30 |
Microscope | Q. Luo, X. Wu, P. Zhao, Y. Nan, W. Chang, X. Zhu, D. Su and Z. Liu,"OTUD1 activates caspase‐independent and caspase‐dependent apoptosis by promoting AIF nuclear translocation and MCL1 degradation", Adv Sci (Weinh)., 2021, doi: 10.1002/advs.202002874. |
| 20 | Cell: Macrophage | Microscope | G. Yang, M. Fan, J. Zhu, C. Ling, L. Wu, X. Zhang, M. Zhang, J. Li, Q. Yao, Z. Gu and X. Cai, "A multifunctional anti-inflammatory drug that can specifically target activated macrophages massively deplete intracellular H2O2 and produce large amounts CO for a highly efficient treatment of osreoarthritis" , Biomaterials, 2020, doi:10.1016/j.biomaterials.2020.120155. |
| 21 | Cell: MDA-MB-415, MCF-7 |
Microscope | S.Y. Park, K.J. Jeong, A. Poire, D. Zhang, Y.H. Tsang, A.S. Blucher and G.B. Mills ,"Irreversible HER2 inhibitors overcome resistance to the RSL3 ferroptosis inducer in non-HER2 amplified luminal breast cancer", Cell Death & Disease., 2023, doi: 10.1038/s41419-023-06042-1. |
| 22 | Cell: MIN6 |
Plate reader/ Microscope |
N. Mizusawa, N. Harada, T. Iwata, I. Ohigashi, M. Itakura and K. Yoshimoto,"Identification of protease serine S1 family member 53 as a mitochondrial protein in murine islet beta cells", Islets., 2022, doi: 10.1080/19382014.2021.1982325. |
| 23 | Cell: MSCs |
Flow Cytometer | S.Y. Jo, H.J. Cho and T.M. Kim,"Fenoldopam mesylate enhances the survival of mesenchymal stem cells under oxidative stress and increases the therapeutic function in acute kidney injury", Cell Transplant., 2023, doi: 10.1177/09636897221147920. |
| 24 | Cell: Neuro-2A |
Microscope、 Plate reader |
Y. Wang, Y. Shinoda, A. Cheng, I. Kawahata and K. Fukunaga,"Epidermal fatty acid-binding protein 5 (FABP5) Involvement in alpha-synuclein-induced mitochondrial injury under oxidative stress", Biomedicines., 2021, doi: 10.3390/biomedicines9020110. |
| 25 | Cell: Neuron |
Microscope | I. Kawahata, L. Luc Bousset, R. Melki and K. Fukunaga , "Fatty Acid-Binding Protein 3 is Critical for α-Synuclein Uptake and MPP+-Induced Mitochondrial Dysfunction in Cultured Dopaminergic Neurons ", Int J Mol Sci., 2019, 20, 5358. |
| 26 | Cell: Neuron |
Microscope | A. Fukuda, S. Nakashima,Y. Oda, K. Nishimura, H. Kawashima, H. Kimura, T. Ohgita, E. Kawashita, K. Ishihara, A. Hanaki, M. Okazaki, E. Matsuda, Y. Tanaka, S. Nakamura, T. Matsumoto, S. Akiba, H. Saito, H. Matsuda and K. Takata,"Plantainoside B in Bacopa monniera Binds to Aβ Aggregates Attenuating Neuronal Damage and Memory Deficits Induced by Aβ", Biol Pharm Bull., 2023, doi: 10.1248/bpb.b22-00797. |
| 27 | Cell: PAECs |
Plate reader | T. Sakai, H. Takagaki, N. Yamagiwa, M. Ui, S. Hatta and J. Imai,"Effects of the cytoplasm and mitochondrial specific hydroxyl radical scavengers TA293 and mitoTA293 in bleomycin-induced pulmonary fibrosis model mice", Antioxidants (Basel)., 2021, doi: 10.3390/antiox10091398. |
| 28 | Cell: PANC-1 |
Plate reader | W.A. Naime, A. Kimishima, A. Setiawan, J.R. Fahim, M.A. Fouad, M.S. Kamel and M. Arai,"Mitochondrial Targeting in an Anti-Austerity Approach Involving Bioactive Metabolites Isolated from the Marine-Derived Fungus Aspergillus sp.", Marine drugs., 2020, doi: 10.3390/md18110555. |
| 29 | Cell: PANC-1, MIAPaca-2 |
Microscope | T. Taniai, Y. Shirai,Y. Shimada, R. Hamura, M. Yanagaki, N. Takada, T. Horiuchi, K. Haruki, K. Furukawa, T. Uwagawa, K. Tsuboi, Y. Okamoto, S. Shimada, S. Tanaka, T. Ohashi and T. Ikegami,"Inhibition of acid ceramidase elicits mitochondrial dysfunction and oxidative stress in pancreatic cancer cells", Cancer Sci., 2021, doi: 10.1111/cas.15123. |
| 30 | Cell: PC |
Flow Cytometer | R. Hamura, Y. Shirai,Y. Shimada, N. Saito, T. Taniai, T. Horiuchi, N. Takada, Y. Kanegae, T. Ikegami, T. Ohashi and K. Yanaga ,"Suppression of lysosomal acid alpha‐glucosidase impacts the modulation of transcription factor EB translocation in pancreatic cancer", Cancer Sci., 2021, doi: 10.1111/cas.14921. |
| 31 | Cell: Porcine oocytes |
Microscope | W. Hu, Y. Zhang, D. Wang, T. Yang, J. Qi, Y. Zhang, H. Jiang, J Zhang, B. Sun and S. Liang,"Iron Overload-Induced Ferroptosis Impairs Porcine Oocyte Maturation and Subsequent Embryonic Developmental Competence in vitro", Front Cell Dev Biol., 2021, doi: 10.3389/fcell.2021.673291. |
| 32 | Cell: Porcine oocytes |
Microscope | Y. Xiao, B. Yuan, W. Hu, J. Qi, H. Jiang, B. Sun, J. Zhang and S. Liang,"Tributyltin Oxide Exposure During in vitro Maturation Disrupts Oocyte Maturation and Subsequent Embryonic Developmental Competence in Pigs", Front Cell Dev Biol., 2021, doi: 10.3389/fcell.2021.683448. |
| 33 | Cell: RGC-5 |
Plate reader | Y. Aoyama, S. Inagaki, K. Aoshima, Y. Iwata, S. Nakamura, H. Hara and M. Shimazawa,"Involvement of endoplasmic reticulum stress in rotenone-induced leber hereditary optic neuropathy model and the discovery of new therapeutic agents", J Pharmacol Sci . .,2021, doi: 10.1016/j.jphs.2021.07.003. |
| 34 | Cell: SAS,HSC-2 |
Plate reader | K. Yamana, J. Inoue, R. Yoshida, J. Sakata, H. Nakashima, H. Arita, S. Kawaguchi, S. Gohara, Y. Nagao, H. Takeshita, M. Maeshiro, R. Liu, Y. Matsuoka, M. Hirayama, K. Kawahara, M. Nagata, A. Hirosue, R. Toya, R. Murakami, Y. Kuwahara, M. Fukumoto and H. Nakayama,"Extracellular vesicles derived from radioresistant oral squamous cell carcinoma cells contribute to the acquisition of radioresistance via the miR‐503‐3p‐BAK axis", J Extracell Vesicles., 2021, doi: 10.1002/jev2.12169. |
| 35 | Cell: SBC-3 |
Flow Cytometer | N. Takahashi, T. Iguchi, M. Kuroda, M. Mishima and Y. Mimaki,"Novel Oleanane-Type Triterpene Glycosides from the Saponaria officinalis L. Seeds and Apoptosis-Inducing Activity via Mitochondria", Int J Mol Sci., 2022, doi: 10.3390/ijms23042047. |
| 36 | Cell: SH-SY5Y |
Microscope | Q. Guo, I. Kawahata, A. Cheng, H. Wang, W. Jia, H. Yoshino and K. Fukunaga,"Fatty acid-binding proteins 3 and 5 are involved in the initiation of mitochondrial damage in ischemic neurons", Redox Biology., 2023, doi: 10.1016/j.redox.2022.102547. |
| 37 | Cell: SiHa |
Microscope | F.F. Gao, J.H. Quan, M.A. Lee, W. Ye, J.M. Yuk, G.H. Cha, I.W. Choi and Y.H. Lee,"Trichomonas vaginalis induces apoptosis via ROS and ER stress response through ER–mitochondria crosstalk in SiHa cells", Parasites &vectors., 2021, doi: 10.1186/s13071-021-05098-2. |
| 38 | Cell: SU-DHL-2 |
Flow Cytometer | Q. Zhao, D. Jiang, X. Sun, Q. Mo, S. Chen, W. Chen, R. Gui and X. Ma,"Biomimetic nanotherapy: core–shell structured nanocomplexes based on the neutrophil membrane for targeted therapy of lymphoma", J Nanobiotechnology., 2021, doi: 10.1186/s12951-021-00922-4. |
| 39 | Cell: THP-1 |
Microscope | W. Zheng, Z. Zhou, Y. Rui, R. Ye, F. Xia, F. Guo, X. Liu, J. Su, M. Lou, and X.F. Yu,"TRAF3 activates STING-mediated suppression of EV-A71 and target of viral evasion", Signal Transduct Target Ther., 2023, doi: 10.1038/s41392-022-01287-2. |
| 40 | Cell: TSM15 |
In Cell Analyzer | M. Honda, F. Shimizu, R. Sato, Y. Mizukami, K. Watanabe, Y. Takeshita, T. Maeda, M. Koga and T. Kanda,"Jo-1 Antibodies From Myositis Induce Complement-Dependent Cytotoxicity and TREM-1 Upregulation in Muscle Endothelial Cells", Neurol Neuroimmunol Neuroinflamm., 2023, doi: 10.1212/NXI.0000000000200116. |
| 41 | Cell: tumor |
Flow Cytometer | H. Wang, X. Rong, G. Zhao, Y. Zhou, Y. Xiao, D. Ma, X. Jin, Y. Wu, Y. Yan, H. Yang, Y. Zhou, M. Qian, C. Niu, X. Hu, D.Q. Li, Q. Liu, Y. Wen, Y.Z. Jiang, C. Zhao and Z.M. Shao ,"The microbial metabolite trimethylamine N-oxide promotes antitumor immunity in triple-negative breast cancer", Cell Metab., 2022, doi: 10.1016/j.cmet.2022.02.010. |
| 42 | Cell: TY10 |
In Cell Analyzer | F. Shimizu, R. Ogawa, Y. Mizukami, K. Watanabe, K. Hara, C. Kadono, T. Takahashi, T. Misu, Y. Takeshita, Y. Sano, M. Fujisawa, T. Maeda, I. Nakashima, K. Fujihara and T. Kanda,"GRP78 antibodies are associated with blood-brain barrier breakdown in anti–myelin oligodendrocyte glycoprotein antibody–associated disorder", Neurol Neuroimmunol Neuroinflamm., 2022, doi: 10.1212/NXI.0000000000001038. |
| 43 | Cell: U2OS, HeLa |
Microscope | T. Namba, "BAP31 regulates mitochondrial function via interaction with Tom40 within ER-mitochondria contact sites ", Sci Adv., 2019, 5, (6), 1386. |
Q & A
-
Q
How many samples can I analyze with each kit?
-
A
・Fluorescence microscope: 35 mm dish x 25 (2 ml/dish)
・Flow cytometer: 100 tests (0.5 ml/test)
・Plate reader: 96 well plate x 5 (100 µl/well)
-
Q
Can PBS be used instead of HBSS to wash cells after JC-1 staining?
-
A
To reduce cell damage, we recommend the use of HBSS; if HBSS is not available, we recommend washing with medium.
-
Q
Can I use serum containing medium?
-
A
Serum containing medium can be used for cell washing and working solution. For fluorescence observation, we recommend Imaging Buffer, but if serum containing medium is used, phenol red-free medium is recommended.
-
Q
Is it possible to fix cells after staining or staining after cell fixation?
-
A
No, fixation after staining and staining after fixation are not possible.
-
Q
The stimulated sample showed an increase (or decrease) in both red and green fluorescence values. How do I interpret the results?
-
A
Compare the red and green fluorescence ratios.
Calculate the ratio of red fluorescence value/green fluorescence value for each of the stimulated sample and control.
The lower the fluorescence ratio, the lower the mitochondrial membrane potential.Reason for evaluating by red/green ratio
Since JC-1 accumulates in cells in a membrane potential-dependent manner, the concentration of JC-1 per cell may vary depending on the cell condition1)2). (The accumulation concentration of JC-1 is different between control and stimulated samples due to the different cell conditions.)
In addition, JC-1 aggregates and shifts its fluorescence from green to red when the mitochondrial membrane potential is high. The amount of this aggregation depends on the degree of membrane potential3), allowing comparison of mitochondrial membrane potential between samples in terms of the red/green ratio.<References>
1) Cossarizza, A. et al., Biochem Biophys Res Commun. 1993, 197(1), 40.
2) Perelman, A. et al., Cell Death and Disease, 2012, 3, e430
3) Smiley, S. T. et al., Proc. Nail. Acad. Sci. 1991, 88, 3671.
Handling and storage condition
| 0-5°C |